Identifying Cellular and Molecular Signatures in Human Neurons That Distinguish Fragile X Syndrome Patients with Divergent EEG Profiles
Xinyu Zhao, PhD
Principal Investigator
Carissa Sirois, PhD
Co-Principal Investigator
University of Wisconsin-Madison
Madison, WI
2021-2023 Grant Funding: $90,000
Summary
Why is it so hard to find the right medications to help people with Fragile X syndrome? Just as Fragile X affects individuals differently, medications do as well. This project aims to bring personalized medicine to Fragile X.
This team will model a person’s drug response with a sampling of their cells. Rather than test drug after drug on an individual, drug screening can be done non-invasively. By screening medicines with this approach, years of trying to find the best drugs could be avoided.
Future Fragile X clinical trials would also be improved by identifying patients that may respond better to drugs being tested.
The Results
The Science
By Xinyu Zhao, PhD
While there have been many clinical trials for individuals with Fragile X syndrome, most have been unsuccessful. One likely reason for this is that Fragile X patients have variable symptoms and severity, and so may have different responses to drugs.
Electroencephalogram (EEG) is a non-invasive method that measures different forms of brain activities. Dr. Craig Erickson at Cincinnati Children’s Hospital has also shown that some Fragile X patients have what is known as “high resting gamma” activity, while others have resting gamma activity that is in the normal range (“low resting gamma”). Dr. Erickson also found that this resting gamma measurement is related to the severity of certain symptoms and responses to certain clinical trial drugs.
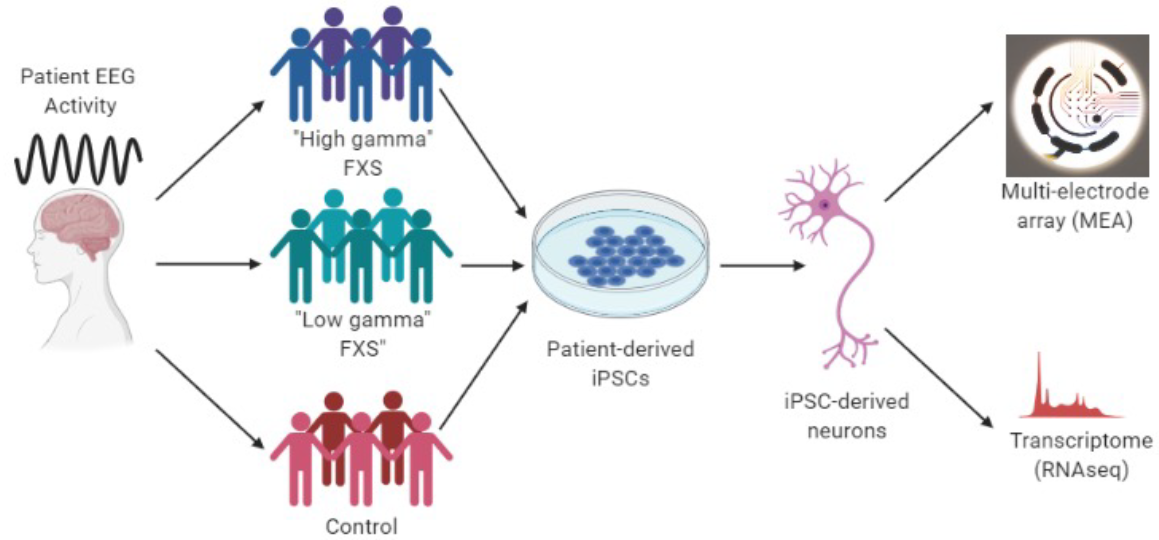
Working closely with Dr. Erickson, we have created induced stem cell lines from the blood cells of Fragile X patients with high resting gamma and Fragile X patients with low resting gamma, as well as unaffected individuals. We have turned these stem cells into human neurons in petri dishes and found that neurons from FXS patients with different EEG exhibit distinct electrical activities. We have developed a high throughput method to detect electrical activities of human neurons in the dish. In this project, we aim to identify electrical signature of neurons from “high gamma” versus “low gamma” Fragile X individuals using a technique called multi-electrode array (MEA). MEA allows us to record from the same neurons over many weeks in culture and can be done in high throughout.
By identifying whether “high gamma” and “low gamma” neurons display unique electrical activity signatures, we can then ask:
- Does this electrical activity signature respond to clinical trial drugs?
- Do “high gamma” and “low gamma” neurons respond to drugs differently?
- Do the neurons in a dish respond to certain drugs the same way that the Fragile X patient did?
Our ultimate goal is to demonstrate that it is possible to model a patient’s drug response in a dish, which could then allow us to screen drug responses in cells. This in turn can help improve clinical trial design by identifying patients that may respond better to certain drugs being tested.